More recently, the new technology of mass cytometry replaces fluorophores with rare earth elements detected by time of flight mass spectrometry, achieving the ability to measure the expression of 34 or more markers.

Through the late 1990s into the mid-2000s, however, rapid development of new fluorophores resulted in modern instruments capable of quantifying up to 18 markers per cell. Until the early 2000s, flow cytometry could only measure a few fluorescent markers at a time. The ability to quantify these has led to flow cytometry being used in a wide range of applications, including but not limited to: These may be endogenous fluorophores such as chlorophyll or transgenic green fluorescent protein, or they may be artificial fluorophores covalently bonded to detection molecules such as antibodies for detecting proteins, or hybridization probes for detecting DNA or RNA. The stream is passed by one or more lasers, and the resulting fluorescent and scattered light is detected by photomultipliers.īy using optical filters, particular fluorophores on or within the cells can be quantified by peaks in their emission spectra. Schematic diagram of a flow cytometer, showing focusing of the fluid sheath, laser, optics (in simplified form, omitting focusing), photomultiplier tubes (PMTs), analogue-to-digital converter, and analysis workstation.įlow cytometers operate by hydrodynamically focusing suspended cells so that they separate from each other within a fluid stream.
#Flowjo license columbia university software
Open software is most widely available in the form of a suite of Bioconductor packages, but is also available for web execution on the GenePattern platform. Open data is slowly growing with the opening of the CytoBank database in 2010, and FlowRepository in 2012, both of which allow users to freely distribute their data, and the latter of which has been recommended as the preferred repository for MIFlowCyt-compliant data by ISAC. Open standards, data and software are also key parts of flow cytometry bioinformatics.ĭata standards include the widely adopted Flow Cytometry Standard (FCS) defining how data from cytometers should be stored, but also several new standards under development by the International Society for Advancement of Cytometry (ISAC) to aid in storing more detailed information about experimental design and analytical steps. It is also possible to characterize data in more comprehensive ways, such as the density-guided binary space partitioning technique known as probability binning, or by combinatorial gating.įinally, diagnosis using flow cytometry data can be aided by supervised learning techniques, and discovery of new cell types of biological importance by high-throughput statistical methods, as part of pipelines incorporating all of the aforementioned methods.
#Flowjo license columbia university manual
For preprocessing, this includes compensating for spectral overlap, transforming data onto scales conducive to visualization and analysis, assessing data for quality, and normalizing data across samples and experiments.įor population identification, tools are available to aid traditional manual identification of populations in two-dimensional scatter plots (gating), to use dimensionality reduction to aid gating, and to find populations automatically in higher dimensional space in a variety of ways. The rapid growth in the multidimensionality and throughput of flow cytometry data, particularly in the 2000s, has led to the creation of a variety of computational analysis methods, data standards, and public databases for the sharing of results.Ĭomputational methods exist to assist in the preprocessing of flow cytometry data, identifying cell populations within it, matching those cell populations across samples, and performing diagnosis and discovery using the results of previous steps.
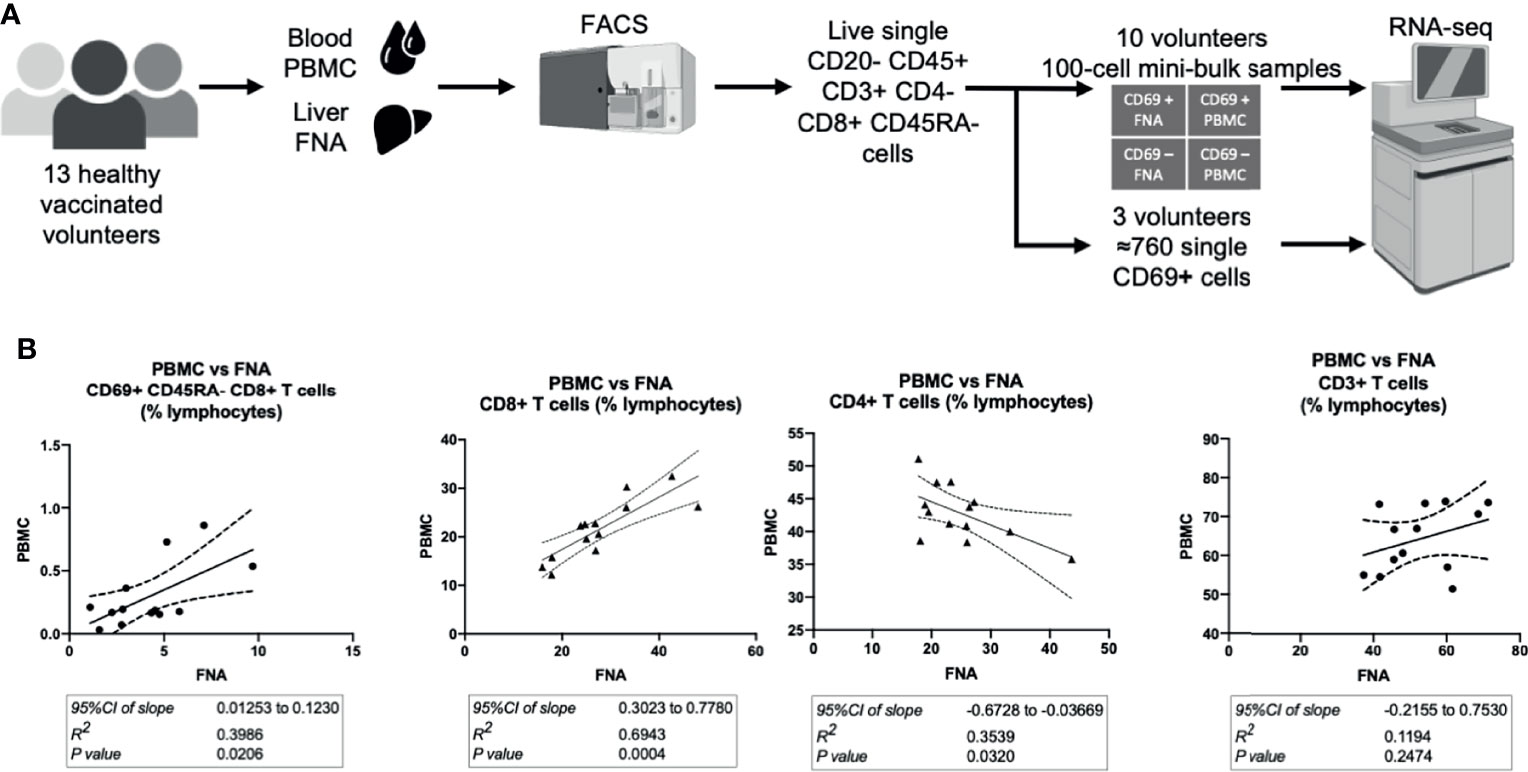
* Corresponding author, email: cytometry bioinformatics is the application of bioinformatics to flow cytometry data, which involves storing, retrieving, organizing and analyzing flow cytometry data using extensive computational resources and tools.įlow cytometry bioinformatics requires extensive use of and contributes to the development of techniques from computational statistics and machine learning.įlow cytometry and related methods allow the quantification of multiple independent biomarkers on large numbers of single cells. Kieran O'Neill 1,2,†, Nima Aghaeepour 1,2,†, Josef Špidlen 1, Ryan Brinkman 1,4*ġTerry Fox Laboratory, BC Cancer Agency, Vancouver, BC, CanadaĢBioinformatics Graduate Program, University of British Columbia, Vancouver, BC, CanadaģDepartment of Medical Genetics, University of British Columbia, British Columbia, Canada
Published under Creative Commons License CC BY 4.0 Public peer review comments can be seen here This article has been published as a PLOS Topic Page
